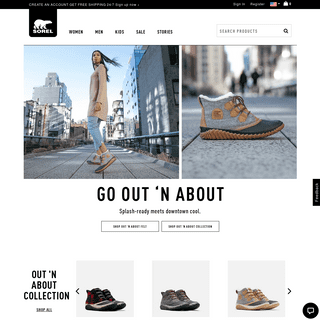
Are you over 18 and want to see adult content?
More Annotations
A complete backup of unprogrammed.org
Are you over 18 and want to see adult content?

A complete backup of aikidoyukishudokan.com
Are you over 18 and want to see adult content?
A complete backup of over40andamumtoone.com
Are you over 18 and want to see adult content?
Favourite Annotations
A complete backup of rajuspot.blogspot.com
Are you over 18 and want to see adult content?
A complete backup of nestreetriders.com
Are you over 18 and want to see adult content?

A complete backup of compartilhamentoespartano.com.br
Are you over 18 and want to see adult content?

A complete backup of iovercome-depression.blogspot.com
Are you over 18 and want to see adult content?
Text
Not logged in
* Log in
Bioinformatics.org
_Membership (41172+) _* Basic Free!
* Professional
_Group hosting _
* Hosted groups (565)_Wiki _
_Franklin Award _
_Sponsorships _
Careers
_About bioinformatics_
_Bioinformatics training _* On-demand courses
* Course bundle
* 10-week program
_Bioinformatics jobs _* Listings
* Employers
Research
_All information groups_
_Online databases_
* EST clusters
* Immigrant genes
* Leukemia genes
* p53 tumor protein gene * Pancreatic cancer genes* TB drug targets
* Acronyms
_Online analysis tools_ * FirstGlance in Jmol * Atlas of Macromolecules * SMS 2: Sequence manipulation * JaMBW: Mol. Biol. workbench * PeCoP: Conserved positions * PrimerX: Mutagenic primers * Savvy: Plasmid map drawing * SeWeR: Sequence analysis * Sequence Extractor _Online education tools_ * Atlas of Macromolecules_More tools _
Development
_All software groups_
_FTP repository _
_SVN & CVS repositories_
_Mailing lists _
Forums
_News & Commentary_ * Submit* Archives
* Subscribe
_Jobs Forum
(Career Center)_ * Submit* Archives
* Subscribe
Latest announcementsSUBMIT
ARCHIVE
SUBSCRIBE
JOURNALS: CFP FOR SPECIAL COLLECTION: PARALLEL COMPUTING IN EVOLUTIONARY BIOINFORMATICS, SAGE JOURNAL Submitted by Jose Luis Guisado Lizar;
posted on Tuesday, July 30, 2019 Special Collection: Parallel Computing in Evolutionary Bioinformatics Journal: Evolutionary Bioinformatics JCR Impact Factor (2018): 2.203 (Q2) Publisher: SAGE Publishing, USA journals.sagepub.com/pageaticsOBJECTIVES
In the last decade, the omics revolution and the application of deep AI technologies to evolutionary bioinformatics have significantly boosted the computing power and storage demands of this field. At the same time, high performance computing (HPC) has also witnessed a revolution that has turned it into a jungle populated by very different types of parallel computing systems exploiting parallelism at different levels. They include multi- and many-core CPUs, GPUs, FPGAs, TPUs or heterogeneous systems joining together several of those types of devices. In addition, cloud computing techniques can also contribute to foster the application of bioinformatics algorithms by turning them into a service (Bioinformatics as a Service, BaaS). The aim of this Special Collection is to present the state of the art on the emerging challenges and achievements regarding the use of parallel computing hardware and software techniques applied to evolutionary bioinformatics and computational evolutionary biology.CALL FOR PAPERS
Accepted papers will be published continuously from Aug. 2019 to Aug31, 2020.
Publication in approximately 4-5 weeks after final acceptance. Topics of interest include, but are not limited to, the application of the following hardware or software techniques to accelerate evolutionary bioinformatics applications: * Multicore and manycore CPUs* Clusters
* Emerging hardware accelerators: GPUs, FPGAs, TPUs* Cloud computing
* Grid computing
* Parallel deep learning * Neuromorphic computing * Unconventional computing techniques ABOUT SPECIAL COLLECTIONS A Special Collection is an opportunity for an open access journal to cultivate a collection of articles around a specific topic, meeting/conference, or a newsworthy development, much in the way a Special Issue functions for a subscription journal. Special Collections are highlighted on the homepage for increased visibility and in most cases receive their own dedicated marketing efforts. Evolutionary Bioinformatics is an open access, peer-reviewed journal. Each article accepted by peer review is made freely available online immediately upon publication, is published under a Creative Commons license and will be hosted online in perpetuity. There is no fee for submitting an article. If, after peer review, your manuscript is accepted for publication, a one-time article processing charge (APC) is payable. Articles invited to submit to a Special Collection are eligible for a discounted Article Processing Charge (APC). Should your article be accepted, you will receive 50% off of the current standing APC, which you can find listed on the journal homepage. MANUSCRIPT PREPARATION AND SUBMISSION Please select the Special Collection title when prompted during the submission process to indicate your interest in publishing in thisSpecial Collection.
Follow the guidelines in the Special Collection web page: journals.sagepub.com/pageatics EVENTS: WEBINAR: ROLE OF LIMS IN OVERCOMING BIOREPOSITORY OPERATIONAL DATA MANAGEMENT CHALLENGESSubmitted by Smith
;
posted on Friday, July 26, 2019 August 13, 2019, 5 PM GMT | 1 PM EDTOnline
cloudlims.com
CloudLIMS, an ISO 9001:2015 certified informatics company, is pleased to host a complimentary webinar titled "Role of LIMS in Overcoming Biorepository Operational Data Management Challenges." The webinar will be presented by Dr. Steven Haynes, the manager of the Sheffield Biorepository, which is a part of the University of Sheffield'sMedical School.
OVERVIEW
This 30-minute webinar iterates the significance of biobanks in research and outlines the regulatory requirements of biobanks such as HIPAA, FDA's 21 CFR Part 11, IRB, audit trail, etc. The webinar also highlights the factors to consider when sourcing biospecimens from biobanks, the challenges involved in managing massive volumes of samples and the associated data, such as demographics, clinical history, and informed consent. Furthermore, the webinar illustrates the role of a SaaS LIMS in the cloud to securely manage data, follow regulatory guidelines, enhance operational efficiency, maximize investment returns, and contribute to the sustainability of a biobank.KEY TAKEAWAYS
* Role of biobanks in research * Data management, quality, regulatory, and operational challengesof biobanks
* Role of a LIMS in addressing such challenges at The University ofSheffield
* Benefits offered by a SaaS LIMS in the Cloud SOFTWARE: ZDNET: IBM GIVES CANCER-KILLING DRUG AI PROJECT TO THE OPENSOURCE COMMUNITY
Submitted by J.W. Bizzaro;
posted on Wednesday, July 24, 2019EXCERPT
> IBM has released three artificial intelligence (AI) projects > tailored to take on the challenge of curing cancer to the > open-source community.>
> At the 18th European Conference on Computational Biology (ECCB) and > the 27th Conference on Intelligent Systems for Molecular Biology > (ISMB), which will be held in Switzerland later this month, the tech > giant will dive into how each of the projects can advance our > understanding of cancers and their treatment. Source: www.zdnet.com/artirugs/AVAILABILITY
github.com/drugcmann github.com/drugeract RESEARCH: UNIVERSITY OF TOKYO: BEYOND FINDING A GENE: SAME REPEATED STRETCH OF DNA IDENTIFIED IN THREE NEURODEGENERATIVE DISEASES Submitted by J.W. Bizzaro;
posted on Monday, July 22, 2019EXCERPT
> Families living with extremely rare neurodegenerative diseases > finally received an answer to the cause of their illnesses, thanks > to a researcher's hunch and decades of improvements in DNA > sequencing technology.>
> Four different rare diseases are all caused by the same short > segment of DNA repeated too many times, a mutation researchers call > noncoding expanded tandem repeats. Researchers suspect variations of > this type of mutation may cause other diseases that have thus far > evaded diagnosis by genetic testing. Source: www.u-tokyo.ac.jp/focu.html EVENTS: CFP (LAST EXTENDED DEADLINE): NEW PERSPECTIVES IN SCIENCE EDUCATION INTERNATIONAL CONFERENCE - 9TH EDITION Submitted by Andrea Baldini;
posted on Thursday, July 18, 2019March 19-20, 2020
Florence, Italy
conference.pixel-online.net/NPSEOBJECTIVES & SCOPE
The objective of the Conference is to promote transnational cooperation and share good practice in the field of innovation for science education. The New Perspectives in Science Education Conference is also an excellent opportunity for the presentation of previous and current projects in the science field. The Call for Papers is addressed to teachers, researchers and experts in the field of science education as well as to coordinators of science and training projects. Experts in the field of science teaching and learning are therefore invited to submit an abstract of a paper to be presented during theconference.
PRESENTATIONS
There will be three presentation modalities: oral, poster and virtualpresentations.
All accepted papers will be included in the Conference Proceedings published by Filodiritto Editore with ISBN and ISSN codes. This publication will be sent to be reviewed for inclusion in Conference Proceedings Citation Index by Thomson Reuters (ISI-Clarivate). The publication will also be included in Academia.edu and indexed inGoogle Scholar.
IMPORTANT DATES
Deadline for submitting abstracts: October 22, 2019 Notification of abstracts' acceptance / rejection: November 5, 2019 Deadline for papers' submission: January 20, 2020 Conference days: March 19-20, 2020 FOR MORE INFORMATION For further information, please contact us at the following address: sciencepixel-online.net or visit the New Perspectives in Science Education conference website. EVENTS: 14TH WORKFLOWS IN SUPPORT OF LARGE-SCALE SCIENCE WORKSHOP(WORKS 2019)
Submitted by Hoang Nguyen;
posted on Thursday, July 18, 2019 Sunday 17 November 2019Denver, CO, USA
works.cs.cardiff.ac.uk Held in conjunction with SC19, sc19.supercomputing.org Extended paper submission deadline: July 29, 2019SCOPE
Data-intensive Workflows (a.k.a. scientific workflows) are routinely used in most scientific disciplines today, especially in the context of parallel and distributed computing. Workflows provide a systematic way of describing the analysis and rely on workflow management systems to execute the complex analyses on a variety of distributed resources. They are at the interface between end-users and computing infrastructures. With the dramatic increase of raw data volume in every domain, they play an even more critical role to assist scientists in organizing and processing their data and to leverage HPC or HTC resources, e.g., workflows played an important role in the discovery of Gravitational Waves. This workshop focuses on the many facets of data-intensive workflow management systems, ranging from job execution to service management and the coordination of data, service and job dependencies. The workshop therefore covers a broad range of issues in the scientific workflow lifecycle that include: data-intensive workflows representation and enactment; designing workflow composition interfaces; workflow mapping techniques that may optimize the execution of the workflow; workflow enactment engines that need to deal with failures in the application and execution environment; and a number of computer science problems related to scientific workflows such as semantic technologies, compiler methods, fault detection andtolerance.
TOPICS
The topics of the workshop include but are not limited to: * Big Data analytics workflows * Data-driven workflow processing (including stream-based workflows) * Workflow composition, tools, and languages * Workflow execution in distributed environments (including HPC,clouds, and grids)
* Reproducible computational research using workflows * Dynamic data dependent workflow systems solutions * Exascale computing with workflows * In Situ Data Analytics Workflows * Interactive workflows (including workflow steering) * Workflow fault-tolerance and recovery techniques * Workflow user environments, including portals * Workflow applications and their requirements * Adaptive workflows * Workflow optimizations (including scheduling and energyefficiency)
* Performance analysis of workflows * Workflow debugging * Workflow provenanceIMPORTANT DATES
Papers due: July 29, 2019 Paper acceptance notification: September 1, 2019 E-copyright registration completed by authors: October 1, 2019 Camera-ready deadline: October 1, 2019PAPER SUBMISSION
Submitted papers must be at most 10 pages long. The proceedings should be formatted according to the IEEE format (see www.ieee.org/conf.html). The
10-page limit includes figures, tables, appendices and references. WORKS papers will be published in cooperation with TCHPC and will be available from IEEE digital repository. SOFTWARE: BIRCH BIOINFORMATICS SYSTEM 3.50 Submitted by Brian Fristensky;
posted on Wednesday, July 17, 2019NEW
Introducing blreads: BioLegato GUI that makes it easy to run programs involving sequencing reads including * Read trimming, error correction and quality control * Genome assembly and quality assessment * Transcriptome assembly and quality assesment * Generation of differential expression data from readsBIRCH is:
* A comprehensive desktop bioinformatics system which comes with many of the commonly-used bioinformatics programs pre-installed * A framework of tools, files, and documentation for organizing and managing a bioinformatics core facility * An expandable system that allows you to merge 3rd party programs and documentation seamlessly into the standard BIRCH distribution Download BIRCH at home.cc.umanitoba.ca/%7Ep.html Visit our YouTube Channel at www.youtube.com/chanublic RESEARCH: UNIVERSITY OF TOKYO: RESEARCHERS CAN FINALLY MODIFY PLANTMITOCHONDRIAL DNA
Submitted by J.W. Bizzaro;
posted on Wednesday, July 10, 2019EXCERPT
> Researchers in Japan have edited plant mitochondrial DNA for the > first time, which could lead to a more secure food supply.>
> Nuclear DNA was first edited in the early 1970s, chloroplast DNA was > first edited in 1988, and animal mitochondrial DNA was edited in > 2008. However, no tool previously successfully edited plant > mitochondrial DNA.>
> Researchers used their technique to create four new lines of rice > and three new lines of rapeseed (canola). Source: www.u-tokyo.ac.jp/focu.html EVENTS: ROCKY MOUNTAIN BIOINFORMATICS CONFERENCE (ROCKY 2019) Submitted by Suzi Smith;
posted on Wednesday, July 10, 2019December 5-7, 2019
Snowmass Village, CO, USA www.iscb.org/rocky2019 The Rocky series began seventeen years ago as a regional conference, and has grown into an international program with a spotlight on regional development in the computational biosciences. The presenters of the Rocky conference are scientists representing a broad spectrum of universities, industrial enterprises, government laboratories, and medical libraries from around the world. The meeting is a chance to get to know your colleagues near and far, seek collaborative opportunities, and find synergies that can drive our field forward. RESOURCES: PRRDB 2.0: A COMPREHENSIVE DATABASE OF PATTERN-RECOGNITION RECEPTORS AND THEIR LIGANDS Submitted by Gajendra P.S. Raghava;
posted on Tuesday, July 09, 2019 I am happy to share the database PRRDB2, developed by our group for pattern recognition receptors. This will be useful for researchers working in the field of innate immunology or for designing vaccineadjuvants.
AVAILABILITY
Website: webs.iiitd.edu.in/raghava/prrdb2/ Paper: doi.org/10.1az076SUBMIT
ARCHIVE
SUBSCRIBE
Acknowledgments
We wish to thank the following for their support: Copyright © 2019 · Scilico, LLCDetails
Copyright © 2024 ArchiveBay.com. All rights reserved. Terms of Use | Privacy Policy | DMCA | 2021 | Feedback | Advertising | RSS 2.0