Are you over 18 and want to see adult content?
More Annotations

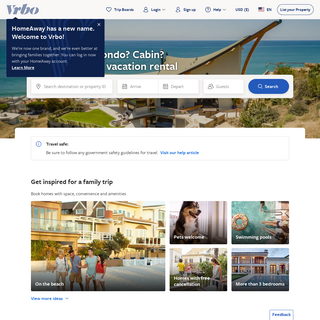
A complete backup of travell-10.blogspot.com
Are you over 18 and want to see adult content?
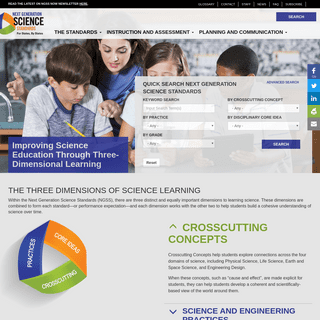
A complete backup of nextgenscience.org
Are you over 18 and want to see adult content?

A complete backup of ringautomotive.com
Are you over 18 and want to see adult content?

A complete backup of auroraclothing.co.nz
Are you over 18 and want to see adult content?
Favourite Annotations
A complete backup of recargapay.com.br
Are you over 18 and want to see adult content?

A complete backup of thegamesjournal.com
Are you over 18 and want to see adult content?
A complete backup of advertisercommunity.com
Are you over 18 and want to see adult content?
A complete backup of incredibleegg.org
Are you over 18 and want to see adult content?

A complete backup of terrorismawareness.org
Are you over 18 and want to see adult content?

A complete backup of esportsobserver.com
Are you over 18 and want to see adult content?
Text
BEAST SOFTWARE
BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. It can be used as a method of reconstructing phylogenies but is also a framework for testing evolutionary hypotheses withoutFIRST TUTORIAL
First Tutorial. An introductory tutorial to getting started with BEAST. This tutorial describes the use of BEAUti and BEAST to analyse some primate sequences and estimate a phylogenetic tree. It will take you through the process of importing an alignment, making choices about the model, generating a BEAST XML file. You will then run BEAST. EFFECTIVE SAMPLE SIZE (ESS) The ESS is calculated by measuring the correlation between sampled states in the chain (i.e., the entries in the log file). If the sampling frequency is very low these will be uncorrelated. This will be indicated by the ESS being approximately equal to the number of states in the log file (minus the burn-in). If this is the case, thenit may be
CONSTRUCT A MODEL IN BEAUTITREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
BEAST V1.10.4 RELEASED �2021 BEAST Developers. All rights reserved. Site uses a modified Documentation Theme for Jekyll by Tom Johnson. Site last generated:Mar 26, 2021
SECOND TUTORIAL
BEAST V1.8.4 RELEASED Download BEAST v1.8.4 binaries for Mac, Windows and UNIX/Linux. Version 1.8.4 released 17th June 2016 New Features: New structured list of citations printed to screen before running. Option ('-citation_file') to write citation list to file.TREE PRIORS
LOGCOMBINER
To run logcombiner on the command-line you can use the shell script installed in /bin of the BEAST package. Use the help command to see a list of options: bin/logcombiner -help. LogCombiner v1.10, 2002-2017 MCMC Output Combiner by Andrew Rambaut and Alexei J. Drummond Institute of Evolutionary Biology University of Edinburgha.rambaut@ed.ac.uk
BEAST SOFTWARE
BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. It can be used as a method of reconstructing phylogenies but is also a framework for testing evolutionary hypotheses withoutFIRST TUTORIAL
First Tutorial. An introductory tutorial to getting started with BEAST. This tutorial describes the use of BEAUti and BEAST to analyse some primate sequences and estimate a phylogenetic tree. It will take you through the process of importing an alignment, making choices about the model, generating a BEAST XML file. You will then run BEAST. EFFECTIVE SAMPLE SIZE (ESS) The ESS is calculated by measuring the correlation between sampled states in the chain (i.e., the entries in the log file). If the sampling frequency is very low these will be uncorrelated. This will be indicated by the ESS being approximately equal to the number of states in the log file (minus the burn-in). If this is the case, thenit may be
CONSTRUCT A MODEL IN BEAUTITREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
BEAST V1.10.4 RELEASED �2021 BEAST Developers. All rights reserved. Site uses a modified Documentation Theme for Jekyll by Tom Johnson. Site last generated:Mar 26, 2021
SECOND TUTORIAL
BEAST V1.8.4 RELEASED Download BEAST v1.8.4 binaries for Mac, Windows and UNIX/Linux. Version 1.8.4 released 17th June 2016 New Features: New structured list of citations printed to screen before running. Option ('-citation_file') to write citation list to file.TREE PRIORS
LOGCOMBINER
To run logcombiner on the command-line you can use the shell script installed in /bin of the BEAST package. Use the help command to see a list of options: bin/logcombiner -help. LogCombiner v1.10, 2002-2017 MCMC Output Combiner by Andrew Rambaut and Alexei J. Drummond Institute of Evolutionary Biology University of Edinburgha.rambaut@ed.ac.uk
INSTALLING BEAST
Installing BEAST. BEAST has been developed in Java, which allows the same code to run on any platform that has the Java software installed.We have also created packages for each of the common operating systems to provide a user-interface that is ‘native’ andfamiliar.
TREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
OPTIMIZING PERFORMANCE The idea then is to put the smallest partition (s) on the CPU and the large (r) partitions on the GPU (s). For example, in the case of a small first partitions and four larger subsequent partitions, we suggest to use the following command: beast -beagle_gpu -beagle_order0,2,3,2,3. or:
BEAUTI & THE BEAST
Bayesian Evolutionary Analysis Utility (BEAUti) BEAUti is a graphical user-interface (GUI) application for generating BEAST XML files. It is distributed in the same package as BEAST. To download it, go to the BEAST download page.MOLECULAR CLOCKS
Molecular Clocks. BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. However, the clock models in BEAST - which are discussed on this page - may also be used when analysing TRACER | BEAST DOCUMENTATION Tracer. Tracer is a graphical tool for visualization and diagnostics of MCMC output. Tracer (now at version 1.7.1) is a software package for visualising and analysing the MCMC trace files generated through Bayesian phylogenetic inference. Tracer provides kernel density estimation, multivariate visualisation, demographic trajectory SPREAD3 | BEAST DOCUMENTATION SpreaD3. SpreaD3 (Spatial Phylogenetics Reconstruction of Evolutionary Dynamics using Data-Driven Documents (D3)) is a major update to SPREAD and constitutes a package for analysis and visualisation of pathogen pylodynamic reconstructions. MODEL SELECTION TUTORIAL Marginal likelihood estimation using path sampling and stepping-stone sampling. Recent years have seen the development of several new approaches to perform model selection in the field of phylogenetics, such as path sampling (under the term ‘thermodynamic integration’; Lartillot and Philippe, 2006), stepping-stone sampling (Xie et al., 2011) and generalized stepping-stone sampling (Fan et HOW TO SPECIFY DATES FOR TIPS A key feature of BEAST is the ability to specify dates for individual sequences (i.e., tips in the tree --- so called tipdates). This is important when the organism being studies is evolving on the same scale as the time range of the tree. Common cases of this are fast evolving pathogens such as viruses and bacteria or DNA recovered fromSUMMARIZING TREES
Approaches to summarizing trees. 50% Majority consensus tree and extended majority consensus tree. The 50% majority consensus tree is a tree constructed so that it contains all of the clades that occur in at least 50% of the trees in the posterior distribution.BEAST SOFTWARE
BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. It can be used as a method of reconstructing phylogenies but is also a framework for testing evolutionary hypotheses withoutTHE BEAST PROGRAM
The BEAST program. This program takes as input an XML command file and returns as output log files. The log files record a sample of the states that the Markov chain encountered. This output must be analyzed to produce estimates of the parameters of interest (evolutionary rates, divergence times, population sizes and tree topologies). CONSTRUCT A MODEL IN BEAUTI EFFECTIVE SAMPLE SIZE (ESS) The ESS is calculated by measuring the correlation between sampled states in the chain (i.e., the entries in the log file). If the sampling frequency is very low these will be uncorrelated. This will be indicated by the ESS being approximately equal to the number of states in the log file (minus the burn-in). If this is the case, thenit may be
TREE PRIORS
TREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
BEAST V1.10.4 RELEASED �2021 BEAST Developers. All rights reserved. Site uses a modified Documentation Theme for Jekyll by Tom Johnson. Site last generated:Mar 26, 2021
LOGCOMBINER
To run logcombiner on the command-line you can use the shell script installed in /bin of the BEAST package. Use the help command to see a list of options: bin/logcombiner -help. LogCombiner v1.10, 2002-2017 MCMC Output Combiner by Andrew Rambaut and Alexei J. Drummond Institute of Evolutionary Biology University of Edinburgha.rambaut@ed.ac.uk
HOW TO SPECIFY DATES FOR TIPS A key feature of BEAST is the ability to specify dates for individual sequences (i.e., tips in the tree --- so called tipdates). This is important when the organism being studies is evolving on the same scale as the time range of the tree. Common cases of this are fast evolving pathogens such as viruses and bacteria or DNA recovered from BEAST V1.8.4 RELEASED Download BEAST v1.8.4 binaries for Mac, Windows and UNIX/Linux. Version 1.8.4 released 17th June 2016 New Features: New structured list of citations printed to screen before running. Option ('-citation_file') to write citation list to file.BEAST SOFTWARE
BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. It can be used as a method of reconstructing phylogenies but is also a framework for testing evolutionary hypotheses withoutTHE BEAST PROGRAM
The BEAST program. This program takes as input an XML command file and returns as output log files. The log files record a sample of the states that the Markov chain encountered. This output must be analyzed to produce estimates of the parameters of interest (evolutionary rates, divergence times, population sizes and tree topologies). CONSTRUCT A MODEL IN BEAUTI EFFECTIVE SAMPLE SIZE (ESS) The ESS is calculated by measuring the correlation between sampled states in the chain (i.e., the entries in the log file). If the sampling frequency is very low these will be uncorrelated. This will be indicated by the ESS being approximately equal to the number of states in the log file (minus the burn-in). If this is the case, thenit may be
TREE PRIORS
TREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
BEAST V1.10.4 RELEASED �2021 BEAST Developers. All rights reserved. Site uses a modified Documentation Theme for Jekyll by Tom Johnson. Site last generated:Mar 26, 2021
LOGCOMBINER
To run logcombiner on the command-line you can use the shell script installed in /bin of the BEAST package. Use the help command to see a list of options: bin/logcombiner -help. LogCombiner v1.10, 2002-2017 MCMC Output Combiner by Andrew Rambaut and Alexei J. Drummond Institute of Evolutionary Biology University of Edinburgha.rambaut@ed.ac.uk
HOW TO SPECIFY DATES FOR TIPS A key feature of BEAST is the ability to specify dates for individual sequences (i.e., tips in the tree --- so called tipdates). This is important when the organism being studies is evolving on the same scale as the time range of the tree. Common cases of this are fast evolving pathogens such as viruses and bacteria or DNA recovered from BEAST V1.8.4 RELEASED Download BEAST v1.8.4 binaries for Mac, Windows and UNIX/Linux. Version 1.8.4 released 17th June 2016 New Features: New structured list of citations printed to screen before running. Option ('-citation_file') to write citation list to file.INSTALLING BEAST
Installing BEAST. BEAST has been developed in Java, which allows the same code to run on any platform that has the Java software installed.We have also created packages for each of the common operating systems to provide a user-interface that is ‘native’ andfamiliar.
THE BEAST PROGRAM
The BEAST program. This program takes as input an XML command file and returns as output log files. The log files record a sample of the states that the Markov chain encountered. This output must be analyzed to produce estimates of the parameters of interest (evolutionary rates, divergence times, population sizes and tree topologies). BEAUTI & THE BEAST AND OTHER PROGRAMS This is the main program that takes a control file generated by BEAUti and performs the analysis. TreeAnnotator | This is a post-analysis program that will produce a summary tree from the output of BEAST. LogCombiner | This is a utility program that will combine log files from different runs and reduce the sampling frequency (thin them).FIRST TUTORIAL
First Tutorial. An introductory tutorial to getting started with BEAST. This tutorial describes the use of BEAUti and BEAST to analyse some primate sequences and estimate a phylogenetic tree. It will take you through the process of importing an alignment, making choices about the model, generating a BEAST XML file. You will then run BEAST.TREEANNOTATOR
TreeAnnotator is a program to summarize the information from a sample of trees produced by BEAST onto a single “target” tree. The summary information includes the posterior probabilities of the nodes in the target tree, the posterior estimates and HPD limits of the node heights and (in the case of a relaxed molecular clock model) therates.
BEAGLE | BEAST DOCUMENTATION To test the installation, run BEAST and when the options dialog box appears, select “Use BEAGLE library” and “Show list of available BEAGLE resources”: The BEAGLE options available in the BEAST dialog box. You don’t need to specify a BEAST input file as the information about BEAGLE will beBEAUTI & THE BEAST
Bayesian Evolutionary Analysis Utility (BEAUti) BEAUti is a graphical user-interface (GUI) application for generating BEAST XML files. It is distributed in the same package as BEAST. To download it, go to the BEAST download page.MOLECULAR CLOCKS
Molecular Clocks. BEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. However, the clock models in BEAST - which are discussed on this page - may also be used when analysingLOGCOMBINER
LogCombiner v1.10, 2002-2017 MCMC Output Combiner by Andrew Rambaut and Alexei J. Drummond Institute of Evolutionary Biology University of Edinburgh a.rambaut@ed.ac.uk Department of Computer Science University of Auckland alexei@cs.auckland.ac.nz Usage: logcombiner[ * * Contents
* FAQ
* Help
* Source Code
* News
*
*
Details
Copyright © 2024 ArchiveBay.com. All rights reserved. Terms of Use | Privacy Policy | DMCA | 2021 | Feedback | Advertising | RSS 2.0